File:Koronavirustapauksia suomessa ennuste muiden maiden datoista 1.svg

Original file (SVG file, nominally 1,058 × 518 pixels, file size: 50 KB)
Captions
Captions
Summary
edit| DescriptionKoronavirustapauksia suomessa ennuste muiden maiden datoista 1.svg |
Suomi: Koronavirustapauksia suomessa ennuste muiden maiden datoista. |
| Date | |
| Source | Own work |
| Author | Merikanto |
| SVG development InfoField |
Python 3 code to produce forecast graph
Increasing trend forecast
-
- COVID-19 Finland short-term forecast
- based on data from other countries
- Python 3 script
-
- Input from internet site: cases, recovered, deaths.
-
- version 0001.0003
- 12.1.2021
-
import math as math
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import locale
from datetime import datetime, timedelta
import matplotlib.dates as mdates
from scipy import interpolate
import scipy.signal
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter,
AutoMinorLocator, MaxNLocator)
from scipy.signal import savgol_filter
- ggplot
- from plotnine import *
from mizani.breaks import date_breaks, minor_breaks
from mizani.formatters import date_format
from bs4 import BeautifulSoup
import requests
import locale
from datetime import datetime, timedelta
import matplotlib.dates as mdates
import matplotlib.ticker as ticker
import matplotlib.dates as mdates
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter,
AutoMinorLocator)
from matplotlib.ticker import MaxNLocator
from matplotlib.ticker import ScalarFormatter
from statsmodels.tsa.statespace.sarimax import SARIMAX
from statsmodels.tsa.arima.model import ARIMA
from numpy import fft
import statistics
from scipy.interpolate import UnivariateSpline
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
from sklearn.ensemble import RandomForestRegressor
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn import svm, metrics
from pygam import LogisticGAM
from pygam import LinearGAM
from sklearn.neighbors import KernelDensity
from scipy.interpolate import pchip_interpolate
from scipy import stats
from datetime import date
from scipy.optimize import curve_fit
- asetukset parameters
- pohjadatan rajat: limits of base data
- paiva1="2021-01-01"
- paiva2="2021-03-19"
- ennusteen rajat: forecast limits
- taika1="2021-03-19"
- taika2="2021-06-01"
paiva1="2021-01-01"
paiva2="2021-07-12"
- ennusteen rajat: forecast limits
taika1="2021-07-12"
taika2="2021-09-01"
- y-akselin rajat
ymax1=1200
ymax2=20
vormat1='%Y-%m-%d'
today = date.today()
- print("Today's date:", today)
yesterday = datetime.today() - timedelta(days=1)
locale.setlocale(locale.LC_TIME, "fi_FI")
def func(x, a, b):
return (a*x+b)
def fourierExtrapolation(x, n_predict):
n = x.size
n_harm = 10 # number of harmonics in model
t = np.arange(0, n)
p = np.polyfit(t, x, 1) # find linear trend in x
x_notrend = x - p[0] * t # detrended x
x_freqdom = fft.fft(x_notrend) # detrended x in frequency domain
f = fft.fftfreq(n) # frequencies
indexes = range(n)
# sort indexes by frequency, lower -> higher
#indexes.sort(key = lambda i: np.absolute(f[i]))
t = np.arange(0, n + n_predict)
restored_sig = np.zeros(t.size)
for i in indexes[:1 + n_harm * 2]:
ampli = np.absolute(x_freqdom[i]) / n # amplitude
phase = np.angle(x_freqdom[i]) # phase
restored_sig += ampli * np.cos(2 * np.pi * f[i] * t + phase)
return restored_sig + p[0] * t
def cut_by_dates(dfx, start_date, end_date):
mask = (dfx['Date'] > start_date) & (dfx['Date'] <= end_date)
dfx2 = dfx.loc[mask]
#print(dfx2)
return(dfx2)
def load_country_cases(maa):
dfin = pd.read_csv('https://datahub.io/core/covid-19/r/countries-aggregated.csv', parse_dates=['Date'])
countries = [maa]
dfin = dfin[dfin['Country'].isin(countries)]
#print (head(dfin))
#quit(-1)
selected_columns = dfin"Date", "Confirmed", "Recovered", "Deaths"
df2 = selected_columns.copy()
df=df2
len1=len(df["Date"])
aktiv2= [None] * len1
for n in range(0,len1-1):
aktiv2[n]=0
dates=df['Date']
rekov1=df['Recovered']
konf1=df['Confirmed']
death1=df['Deaths']
#print(dates)
spanni=6
#print(rekov1)
#quit(-1)
rulla = rekov1.rolling(window=spanni).mean()
rulla2 = rulla.rolling(window=spanni).mean()
tulosrulla=rulla2
tulosrulla= tulosrulla.replace(np.nan, 0)
tulosrulla=np.array(tulosrulla).astype(int)
rulla2=tulosrulla
x=np.linspace(0,len1,len1);
#print("kupla")
#print(tulosrulla)
#print(konf1)
#print(death1)
#print(aktiv2)
konf1=np.array(konf1).astype(int)
death1=np.array(death1).astype(int)
#print(konf1)
#quit(-1)
for n in range(0,(len1-1)):
#print("luzmu")
rulla2[n]=tulosrulla[n]
#print ("luzmu2")
#aktiv2[n]=konf1[n]-death1[n]-rulla2[n]
aktiv2[n]=konf1[n]
#print(rulla2[n])
#quit(-1)
#aktiv3=np.array(aktiv2).astype(int)
dailycases1= [0] * len1
dailydeaths1= [0] * len1
for n in range(1,(len1-1)):
dailycases1[n]=konf1[n]-konf1[n-1]
if (dailycases1[n]<0): dailycases1[n]=0
for n in range(1,(len1-1)):
dailydeaths1[n]=death1[n]-death1[n-1]
if (dailydeaths1[n]<0): dailydeaths1[n]=0
#quit(-1)
df.insert (2, "Daily_Cases", dailycases1)
df.insert (3, "Daily_Deaths", dailydeaths1)
df['ActiveEst']=aktiv2
#print (df)
dfout = df'Date', 'Confirmed','Deaths','Recovered', 'ActiveEst','Daily_Cases','Daily_Deaths'
#print(df)
#print(dfout)
print(".")
return(dfout)
def load_fin_wiki_data():
url="https://fi.wikipedia.org/wiki/Suomen_koronaviruspandemian_aikajana"
response = requests.get(url)
soup = BeautifulSoup(response.text, 'lxml')
table = soup.find_all('table')[0] # Grab the first table
df = pd.read_html(str(table))[0]
#print(df)
#Päivä Tapauksia Uusia tapauksia Sairaalassa Teholla Kuolleita Uusia kuolleita Toipuneita
df2 = df'Tapauksia','Uusia tapauksia','Sairaalassa','Teholla','Kuolleita','Uusia kuolleita','Toipuneita'
kaikkiatapauksia=df['Tapauksia']
toipuneita=df['Toipuneita']
uusiatapauksia=df['Uusia tapauksia']
sairaalassa=df['Sairaalassa']
teholla=df['Teholla']
kuolleita=df['Kuolleita']
uusiakuolleita=df['Uusia kuolleita']
len1=len(kaikkiatapauksia)
kaikkiatapauksia2=[]
toipuneita2=[]
uusiatapauksia2=[]
sairaalassa2=[]
teholla2=[]
kuolleita2=[]
uusiakuolleita2=[]
for n in range(0,len1):
elem0=kaikkiatapauksia[n]
elem1 = .join(c for c in elem0 if c.isdigit())
elem2=int(elem1)
kaikkiatapauksia2.append(elem2)
elem0=toipuneita[n]
elem1 = .join(c for c in elem0 if c.isdigit())
toipuneita2.append(int(elem1))
elem0=uusiatapauksia[n]
elem1 = .join(c for c in elem0 if c.isdigit())
uusiatapauksia2.append(int(elem1))
elem0=sairaalassa[n]
#elem1 = .join(c for c in elem0 if c.isdigit())
sairaalassa2.append(int(elem0))
elem0=teholla[n]
#elem1 = .join(c for c in elem0 if c.isdigit())
teholla2.append(int(elem0))
elem0=kuolleita[n]
#elem1 = .join(c for c in elem0 if c.isdigit())
kuolleita2.append(int(elem0))
elem0=uusiakuolleita[n]
#elem1 = .join(c for c in elem0 if c.isdigit())
uusiakuolleita2.append(int(elem0))
#kaikkiatapauksia3=np.array(kaikkiatapauksia2).astype(int)
#print("---")
#print(kaikkiatapauksia2)
#print(toipuneita2)
kaikkiatapauksia3=np.array(kaikkiatapauksia2).astype(int)
toipuneita3=np.array(toipuneita2).astype(int)
uusiatapauksia3=np.array(uusiatapauksia2).astype(int)
sairaalassa3=np.array(sairaalassa2).astype(int)
teholla3=np.array(teholla2).astype(int)
kuolleita3=np.array(kuolleita2) .astype(int)
uusiakuolleita3=np.array(uusiakuolleita2).astype(int)
napapaiva1 = np.datetime64("2020-04-01")
timedelta1= np.timedelta64(len(kaikkiatapauksia3),'D')
napapaiva2 = napapaiva1+timedelta1
#dada1 = np.linspace(napapaiva1.astype('f8'), napapaiva2.astype('f8'), dtype='<M8[D]')
dada1 = pd.date_range(napapaiva1, napapaiva2, periods=len(kaikkiatapauksia3)).to_pydatetime()
#print(dada1)
data = {'Date':dada1,
'Kaikkia tapauksia':kaikkiatapauksia3,
"Uusia tapauksia":uusiatapauksia3,
"Sairaalassa":sairaalassa3,
"Teholla":teholla3,
"Kuolleita":kuolleita3,
"Uusiakuolleita":uusiakuolleita3,
"Toipuneita":toipuneita3
}
df2 = pd.DataFrame(data)
#print(kaikkiatapauksia3)
#print ("Fin wiki data.")
return(df2)
def catch_forecasting_data(toka, last1):
lentoka=len(toka)-2
jono=[]
initu=lentoka
for m in range(0,10):
q=0
for n in range(initu,0, -1):
cc=toka[n]
jono.append(cc)
if (cc<last1):
initu=n-1
if(q==0):
jono.pop()
break
arra00=np.array(jono).astype(int)
arra=arra00[::-1]
return(arra)
-
-
- df=load_country_cases('Finland')
- df2=cut_by_dates(df, paiva1,paiva2)
- countries=["Finland", "Denmark", "Israel", "Sweden"]
- countries=["Finland", "Denmark", "Israel", "France", "Germany", "Italy", "Belgium", "Netherlands", "Portugal", "Austria", "Brazil", "Lithuania", "Mexico", "Argentina", "Latvia"]
countries=["Finland", "Denmark", "Germany"]
num_countries=len(countries)
dfs=[]
for n in range(0,num_countries):
#print (countries[n])
dfs.append(load_country_cases(countries[n]))
- for n in range(0,num_countries):
- print(dfs[n])
dada=[]
for n in range(0,num_countries):
df=dfs[n]
#print(df)
taily=df['Daily_Cases']
#print(taily)
taily2=np.array(taily).astype(int)
dada.append(taily2)
- for n in range(0,num_countries):
- print(dada[n])
- last cases num of "Finland" data
eka=dada[0]
numu=30
last1=eka[len(eka)-2]
arras=[]
for n in range(1,num_countries):
toka=dada[n]
arra=catch_forecasting_data(toka, last1)
arras.append(arra)
- print(arra)
- quit(-1)
maaras=num_countries-1
parras=[]
for n in range(0,maaras):
arra=arras[n]
parra=arra[0:numu]
#print (parra)
parras.append(parra)
supercache=[]
for m in range(0,numu):
cache=[]
for n in range(0,maaras):
parra=parras[n]
print(parra)
mmm=parra[m]
cache.append(mmm)
supercache.append(np.array(cache).astype(int))
average00=[]
deviation00=[]
min00=[]
max00=[]
for n in range(0,numu):
average00.append(statistics.mean(supercache[n]))
deviation00.append(statistics.stdev(supercache[n]))
min00.append(np.min(supercache[n]))
max00.append(np.max(supercache[n]))
average1=np.array(average00).astype(int)
deviation1=np.array(deviation00).astype(int)
min1=np.array(min00).astype(int)
max1=np.array(max00).astype(int)
high1=average1+deviation1/2
low1=average1-deviation1/2
- ystdev=statistics.stdev(ycasesminusavgol)
- mean=statistics.mean(ysavgol)
- print (parra)
vormat1='%Y-%m-%d'
today = date.today()
- print("Today's date:", today)
yesterday = datetime.today() - timedelta(days=1)
timedelta1= np.timedelta64(numu,'D')
beginday=today
endday=beginday+timedelta(numu)
days1 = pd.date_range(beginday, endday, periods=numu).to_pydatetime()
- print (parra)
dateformat1 = mdates.DateFormatter('%d.%m')
plt.xticks(fontsize=16)
plt.yticks(fontsize=16, rotation=0)
ax1=plt.gca()
plt.title("Koronavirustapauksia - ennuste", fontsize=18)
ax1.set_xlabel('Pvm', color='g',size=18)
ax1.set_ylabel('Koronatapauksia', color='#800000',size=18)
ax1.xaxis.set_major_formatter(dateformat1)
for n in range(0,maaras):
plt.plot(days1, parras[n],color='#800000', alpha=0.1 )
plt.plot(days1, average1,color='#F00', linewidth=3)
plt.fill_between(days1, max1, min1, fc='#FF3F3F', alpha=0.25)
for n in range(0,6):
high1=average1+(2*deviation1)/(n+1)
low1=average1-(2*deviation1)/(n+1)
plt.fill_between(days1, high1, low1, fc='#FF3F3F', alpha=n*0.05)
for n in range(0,6):
high1=average1+(deviation1)/(n+1)
low1=average1-(deviation1)/(n+1)
plt.fill_between(days1, high1, low1, fc='#FF1F1F', alpha=n*0.05)
- plt.plot(days1, parras[n],color='#800000', alpha=0.1 )
plt.ylim(0,6000)
plt.show()
- print (last1)
Decreasing trend forecast, 40 days
-
- covid forecast test program
- based on data from other countries
- python 3 script
-
- NOTE: trend must be decreasing
-
- Python 3 script
-
- Input from internet site: cases, recovered, deaths.
-
- version 0000.0002
- 3.6.2021
-
import math as math
import numpy as np
import pandas as pd
import statistics
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import matplotlib.dates as mdates
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter,
AutoMinorLocator, MaxNLocator,ScalarFormatter)
from mizani.breaks import date_breaks, minor_breaks
from mizani.formatters import date_format
from datetime import datetime, timedelta, date
import scipy.signal
from scipy.signal import savgol_filter
from scipy.signal import find_peaks
from bs4 import BeautifulSoup
import requests
import locale
from country_list import countries_for_language
- asetukset parameters
- current covid cases in a day
currentcases=145
- pohjadatan rajat: limits of base data
paiva1="2020-03-01"
paiva2="2021-05-20"
- y-akselin rajat
- ymax1=1200
- ymax2=20
vormat1='%Y-%m-%d'
today = date.today() - timedelta(days=2)
- print("Today's date:", today)
yesterday = datetime.today() - timedelta(days=1)
locale.setlocale(locale.LC_TIME, "fi_FI")
def cut_by_dates(dfx, start_date, end_date):
mask = (dfx['Date'] > start_date) & (dfx['Date'] <= end_date)
dfx2 = dfx.loc[mask]
#print(dfx2)
return(dfx2)
def load_country_cases(maa):
dfin = pd.read_csv('https://datahub.io/core/covid-19/r/countries-aggregated.csv', parse_dates=['Date'])
countries = [maa]
dfin = dfin[dfin['Country'].isin(countries)]
#print (head(dfin))
#quit(-1)
selected_columns = dfin"Date", "Confirmed", "Recovered", "Deaths"
df2 = selected_columns.copy()
df=df2
len1=len(df["Date"])
aktiv2= [None] * len1
for n in range(0,len1-1):
aktiv2[n]=0
dates=df['Date']
rekov1=df['Recovered']
konf1=df['Confirmed']
death1=df['Deaths']
#print(dates)
spanni=6
#print(rekov1)
#quit(-1)
rulla = rekov1.rolling(window=spanni).mean()
rulla2 = rulla.rolling(window=spanni).mean()
tulosrulla=rulla2
tulosrulla= tulosrulla.replace(np.nan, 0)
tulosrulla=np.array(tulosrulla).astype(int)
rulla2=tulosrulla
x=np.linspace(0,len1,len1);
#print("kupla")
#print(tulosrulla)
#print(konf1)
#print(death1)
#print(aktiv2)
konf1=np.array(konf1).astype(int)
death1=np.array(death1).astype(int)
#print(konf1)
#quit(-1)
for n in range(0,(len1-1)):
#print("luzmu")
rulla2[n]=tulosrulla[n]
#print ("luzmu2")
#aktiv2[n]=konf1[n]-death1[n]-rulla2[n]
aktiv2[n]=konf1[n]
#print(rulla2[n])
#quit(-1)
#aktiv3=np.array(aktiv2).astype(int)
dailycases1= [0] * len1
dailydeaths1= [0] * len1
for n in range(1,(len1-1)):
dailycases1[n]=konf1[n]-konf1[n-1]
if (dailycases1[n]<0): dailycases1[n]=0
for n in range(1,(len1-1)):
dailydeaths1[n]=death1[n]-death1[n-1]
if (dailydeaths1[n]<0): dailydeaths1[n]=0
#quit(-1)
df.insert (2, "Daily_Cases", dailycases1)
df.insert (3, "Daily_Deaths", dailydeaths1)
df['ActiveEst']=aktiv2
#print (df)
dfout = df'Date', 'Confirmed','Deaths','Recovered', 'ActiveEst','Daily_Cases','Daily_Deaths'
#print(df)
#print(dfout)
print(".")
return(dfout)
def find_curve_from_peak(leka):
lenleka1=len(leka)
if(lenleka1==0):
taffel=np.zeros(99).astype(int)
return (taffel)
last1=leka[len(leka)-2]
#print(" Peaks ... ")
ekay=np.array(leka).astype(int)
ekasmoothedy = savgol_filter(ekay,45, 3)
tokasmoothedy0=savgol_filter(ekasmoothedy,15, 3)
tokasmoothedy=np.array(tokasmoothedy0).astype(int)
ekapeaks = find_peaks(tokasmoothedy, distance=90 )
#print (ekapeaks)
#print("P1")
ekapeaksx=ekapeaks[0]
ekapeaks1=ekapeaks[1]
#ekapeaks2=ekapeaks[2]
ekapeaksy = [ekasmoothedy[index] for index in ekapeaksx]
ekaalen1=len(tokasmoothedy)
numpeaks1=len(ekapeaksx)
if(numpeaks1==0):
taffel=np.zeros(99).astype(int)
return (taffel)
#taffel=np.zeros(ekaalen1).astype(int)
taffel0=[]
nn=0
eleni1=ekaalen1-1
for m in range(0,1):
#pe=ekapeaksx[m+1]
pe=ekapeaksx[m]
#print (pe, ekaalen1)
edc=1000000
nn=0
for n in range(pe,eleni1):
c=tokasmoothedy[n]
#print("N",n,"c", c)
deltac=edc-c
deltadc=deltac-2
if (deltadc>0):
#taffel[nn]=int(c)
taffel0.append(int(c))
edc=c
nn=nn+1
taffel=np.array(taffel0).astype(int)
#plt.plot(taffel)
#plt.show()
paluu=taffel
return(paluu)
-
-
-
-
- df=load_country_cases('Finland')
- df2=cut_by_dates(df, paiva1,paiva2)
- countries=["Finland", "Denmark", "Israel", "Sweden"]
- countries=["Finland", "Denmark", "Israel", "France", "Germany", "Italy", "Belgium", "Netherlands", "Portugal", "Austria", "Brazil", "Lithuania", "Mexico", "Argentina", "Latvia"]
- countries=["Finland", "Denmark", "Israel", "France", "Germany", "Italy", "Belgium", "Netherlands", "Portugal", "Austria", "Brazil", "Lithuania", "Mexico", "Argentina", "Latvia"]
- countries=["Finland", "Denmark", "Israel"]
- countries=["Finland", "Israel", "Norway", "Denmark"]
- countries=["Finland"]
kountries = dict(countries_for_language('en'))
lenkountries=len(kountries)-1
- print(kountries)
print(lenkountries)
- for n in range(0,lenkountries):
- kountri= kountries.values()[n]
- print(kountri)
countries=[]
maxcountries=lenkountries
- maxcountries=8
nnn=0
for x in enumerate(kountries.items()):
kountry=x[1][1]
countries.append(kountry)
nnn=nnn+1
if(nnn>maxcountries):
break
num_countries=len(countries)
print (countries)
- quit(-1)
dfs=[]
for n in range(0,num_countries):
print (countries[n])
dfs.append(load_country_cases(countries[n]))
- for n in range(0,num_countries):
- print(dfs[n])
dada=[]
for n in range(0,num_countries):
df=dfs[n]
#print(df)
taily=df['Daily_Cases']
#print(taily)
taily2=np.array(taily).astype(int)
dada.append(taily2)
- for n in range(0,num_countries):
- print(dada[n])
- cases num of "Finland" data
eka=dada[0]
multitab=[]
num_countries2=0
for n in range(0,num_countries):
bufu0=dada[n]
taffel=find_curve_from_peak(bufu0)
if(np.max(taffel)==0):
print("Nuls.")
else:
multitab.append(taffel)
num_countries2=num_countries2+1
num_countries2=num_countries2-1
many0=[]
nummany1=0
itemnum=40
for n in range(0,num_countries2):
bufu0=np.array(multitab[n]).astype(float)
maxi=np.max(bufu0).astype(float)
skaled0=bufu0/maxi
lennu1=len(skaled0)
if(lennu1>itemnum):
skaled1=skaled0[0:itemnum]
many0.append(skaled1)
nummany1=nummany1+1
- for n in range(0,nummany1):
#yy=many0[n]
#yys.append(yy)
#plt.plot(yy)
smoothed0=savgol_filter(eka,15, 3)
lensmoothed0=len(smoothed0)-1
- currentsmoothedcases=smoothed0[lensmoothed0-5]
currentsmoothedcases=currentcases
- print(eka)
- print(currentsmoothedcases)
today = date.today()
- print("Today's date:", today)
yesterday = datetime.today() - timedelta(days=1)
timedelta1= np.timedelta64(itemnum,'D')
beginday=today
endday=beginday+timedelta(itemnum)
days1 = pd.date_range(beginday, endday, periods=itemnum).to_pydatetime()
- print (parra)
dateformat1 = mdates.DateFormatter('%d.%m')
plt.xticks(fontsize=16)
plt.yticks(fontsize=16, rotation=0)
ax1=plt.gca()
plt.title("Koronavirustapauksia - ennuste muiden maiden datoista", fontsize=18)
ax1.set_xlabel('Pvm', color='g',size=18)
ax1.set_ylabel('Koronatapauksia', color='#800000',size=18)
ax1.xaxis.set_major_formatter(dateformat1)
yys=[]
for n in range(0,nummany1):
bufu0=many0[n]
yy=currentsmoothedcases*bufu0
yys.append(yy)
- plt.plot(days1, yy,color='#800000', alpha=0.7 )
- plt.plot(yy)
print ("Process ...")
print(nummany1)
print(itemnum)
supercache=[]
for m in range(0,itemnum):
cache=[]
for n in range(0,nummany1):
parra=yys[n]
mmm=parra[m]
cache.append(mmm)
supercache.append(np.array(cache).astype(int))
average00=[]
deviation00=[]
min00=[]
max00=[]
for n in range(0,itemnum):
average00.append(statistics.mean(supercache[n]))
deviation00.append(statistics.stdev(supercache[n]))
min00.append(np.min(supercache[n]))
max00.append(np.max(supercache[n]))
average1=np.array(average00).astype(int)
deviation1=np.array(deviation00).astype(int)
min1=np.array(min00).astype(int)
max1=np.array(max00).astype(int)
high1=average1+deviation1/2
low1=average1-deviation1/2
for n in range(0,nummany1):
yy=yys[n]
plt.plot(days1,yy,color='#800000', alpha=0.3)
plt.plot(days1, average1,color='#F00', linewidth=3)
plt.fill_between(days1, max1, min1, fc='#FF3F3F', alpha=0.25)
for n in range(0,6):
high1=average1+(2*deviation1)/(n+1)
low1=average1-(2*deviation1)/(n+1)
plt.fill_between(days1, high1, low1, fc='#FF3F3F', alpha=n*0.05)
for n in range(0,6):
high1=average1+(deviation1)/(n+1)
low1=average1-(deviation1)/(n+1)
plt.fill_between(days1, high1, low1, fc='#FF1F1F', alpha=n*0.05)
plt.show()
print(".")
QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ
OLD CODE, increasing trend
-
- COVID-19 Finland short-term forecast
- based on data from other countries
- Python 3 script
-
- Input from internet site: cases, recovered, deaths.
-
- version 0001.0004
- 23.3.2021
-
import math as math
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import locale
from datetime import datetime, timedelta
import matplotlib.dates as mdates
from scipy import interpolate
import scipy.signal
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter,
AutoMinorLocator, MaxNLocator)
from scipy.signal import savgol_filter
- ggplot
from mizani.breaks import date_breaks, minor_breaks
from mizani.formatters import date_format
from bs4 import BeautifulSoup
import requests
import locale
from datetime import datetime, timedelta
import matplotlib.dates as mdates
import matplotlib.ticker as ticker
import matplotlib.dates as mdates
from matplotlib.ticker import (MultipleLocator, FormatStrFormatter,
AutoMinorLocator)
from matplotlib.ticker import MaxNLocator
from matplotlib.ticker import ScalarFormatter
from statsmodels.tsa.statespace.sarimax import SARIMAX
from statsmodels.tsa.arima.model import ARIMA
from numpy import fft
import statistics
from scipy.interpolate import UnivariateSpline
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
from sklearn.ensemble import RandomForestRegressor
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn import svm, metrics
from pygam import LogisticGAM
from pygam import LinearGAM
from sklearn.neighbors import KernelDensity
from scipy.interpolate import pchip_interpolate
from scipy import stats
from datetime import date
from scipy.optimize import curve_fit
- asetukset parameters
- pohjadatan rajat: limits of base data
paiva1="2020-11-15"
paiva2="2021-01-20"
- ennusteen rajat: forecast limits
taika1="2021-01-20"
taika2="2021-02-20"
- y-akselin rajat
ymax1=1200
ymax2=20
vormat1='%Y-%m-%d'
today = date.today()
- print("Today's date:", today)
yesterday = datetime.today() - timedelta(days=1)
locale.setlocale(locale.LC_TIME, "fi_FI")
def func(x, a, b):
return (a*x+b)
def fourierExtrapolation(x, n_predict):
n = x.size
n_harm = 10 # number of harmonics in model
t = np.arange(0, n)
p = np.polyfit(t, x, 1) # find linear trend in x
x_notrend = x - p[0] * t # detrended x
x_freqdom = fft.fft(x_notrend) # detrended x in frequency domain
f = fft.fftfreq(n) # frequencies
indexes = range(n)
# sort indexes by frequency, lower -> higher
#indexes.sort(key = lambda i: np.absolute(f[i]))
t = np.arange(0, n + n_predict)
restored_sig = np.zeros(t.size)
for i in indexes[:1 + n_harm * 2]:
ampli = np.absolute(x_freqdom[i]) / n # amplitude
phase = np.angle(x_freqdom[i]) # phase
restored_sig += ampli * np.cos(2 * np.pi * f[i] * t + phase)
return restored_sig + p[0] * t
def cut_by_dates(dfx, start_date, end_date):
mask = (dfx['Date'] > start_date) & (dfx['Date'] <= end_date)
dfx2 = dfx.loc[mask]
#print(dfx2)
return(dfx2)
def load_country_cases(maa):
dfin = pd.read_csv('https://datahub.io/core/covid-19/r/countries-aggregated.csv', parse_dates=['Date'])
countries = [maa]
dfin = dfin[dfin['Country'].isin(countries)]
#print (head(dfin))
#quit(-1)
selected_columns = dfin"Date", "Confirmed", "Recovered", "Deaths"
df2 = selected_columns.copy()
df=df2
len1=len(df["Date"])
aktiv2= [None] * len1
for n in range(0,len1-1):
aktiv2[n]=0
dates=df['Date']
rekov1=df['Recovered']
konf1=df['Confirmed']
death1=df['Deaths']
#print(dates)
spanni=6
#print(rekov1)
#quit(-1)
rulla = rekov1.rolling(window=spanni).mean()
rulla2 = rulla.rolling(window=spanni).mean()
tulosrulla=rulla2
tulosrulla= tulosrulla.replace(np.nan, 0)
tulosrulla=np.array(tulosrulla).astype(int)
rulla2=tulosrulla
x=np.linspace(0,len1,len1);
#print("kupla")
#print(tulosrulla)
#print(konf1)
#print(death1)
#print(aktiv2)
konf1=np.array(konf1).astype(int)
death1=np.array(death1).astype(int)
#print(konf1)
#quit(-1)
for n in range(0,(len1-1)):
#print("luzmu")
rulla2[n]=tulosrulla[n]
#print ("luzmu2")
#aktiv2[n]=konf1[n]-death1[n]-rulla2[n]
aktiv2[n]=konf1[n]
#print(rulla2[n])
#quit(-1)
#aktiv3=np.array(aktiv2).astype(int)
dailycases1= [0] * len1
dailydeaths1= [0] * len1
for n in range(1,(len1-1)):
dailycases1[n]=konf1[n]-konf1[n-1]
if (dailycases1[n]<0): dailycases1[n]=0
for n in range(1,(len1-1)):
dailydeaths1[n]=death1[n]-death1[n-1]
if (dailydeaths1[n]<0): dailydeaths1[n]=0
#quit(-1)
df.insert (2, "Daily_Cases", dailycases1)
df.insert (3, "Daily_Deaths", dailydeaths1)
df['ActiveEst']=aktiv2
#print (df)
dfout = df'Date', 'Confirmed','Deaths','Recovered', 'ActiveEst','Daily_Cases','Daily_Deaths'
#print(df)
#print(dfout)
print(".")
return(dfout)
def load_fin_wiki_data():
url="https://fi.wikipedia.org/wiki/Suomen_koronaviruspandemian_aikajana"
response = requests.get(url)
soup = BeautifulSoup(response.text, 'lxml')
table = soup.find_all('table')[0] # Grab the first table
df = pd.read_html(str(table))[0]
#print(df)
#Päivä Tapauksia Uusia tapauksia Sairaalassa Teholla Kuolleita Uusia kuolleita Toipuneita
df2 = df'Tapauksia','Uusia tapauksia','Sairaalassa','Teholla','Kuolleita','Uusia kuolleita','Toipuneita'
kaikkiatapauksia=df['Tapauksia']
toipuneita=df['Toipuneita']
uusiatapauksia=df['Uusia tapauksia']
sairaalassa=df['Sairaalassa']
teholla=df['Teholla']
kuolleita=df['Kuolleita']
uusiakuolleita=df['Uusia kuolleita']
len1=len(kaikkiatapauksia)
kaikkiatapauksia2=[]
toipuneita2=[]
uusiatapauksia2=[]
sairaalassa2=[]
teholla2=[]
kuolleita2=[]
uusiakuolleita2=[]
for n in range(0,len1):
elem0=kaikkiatapauksia[n]
elem1 = .join(c for c in elem0 if c.isdigit())
elem2=int(elem1)
kaikkiatapauksia2.append(elem2)
elem0=toipuneita[n]
elem1 = .join(c for c in elem0 if c.isdigit())
toipuneita2.append(int(elem1))
elem0=uusiatapauksia[n]
elem1 = .join(c for c in elem0 if c.isdigit())
uusiatapauksia2.append(int(elem1))
elem0=sairaalassa[n]
#elem1 = .join(c for c in elem0 if c.isdigit())
sairaalassa2.append(int(elem0))
elem0=teholla[n]
#elem1 = .join(c for c in elem0 if c.isdigit())
teholla2.append(int(elem0))
elem0=kuolleita[n]
#elem1 = .join(c for c in elem0 if c.isdigit())
kuolleita2.append(int(elem0))
elem0=uusiakuolleita[n]
#elem1 = .join(c for c in elem0 if c.isdigit())
uusiakuolleita2.append(int(elem0))
#kaikkiatapauksia3=np.array(kaikkiatapauksia2).astype(int)
#print("---")
#print(kaikkiatapauksia2)
#print(toipuneita2)
kaikkiatapauksia3=np.array(kaikkiatapauksia2).astype(int)
toipuneita3=np.array(toipuneita2).astype(int)
uusiatapauksia3=np.array(uusiatapauksia2).astype(int)
sairaalassa3=np.array(sairaalassa2).astype(int)
teholla3=np.array(teholla2).astype(int)
kuolleita3=np.array(kuolleita2) .astype(int)
uusiakuolleita3=np.array(uusiakuolleita2).astype(int)
napapaiva1 = np.datetime64("2020-04-01")
timedelta1= np.timedelta64(len(kaikkiatapauksia3),'D')
napapaiva2 = napapaiva1+timedelta1
#dada1 = np.linspace(napapaiva1.astype('f8'), napapaiva2.astype('f8'), dtype='<M8[D]')
dada1 = pd.date_range(napapaiva1, napapaiva2, periods=len(kaikkiatapauksia3)).to_pydatetime()
#print(dada1)
data = {'Date':dada1,
'Kaikkia tapauksia':kaikkiatapauksia3,
"Uusia tapauksia":uusiatapauksia3,
"Sairaalassa":sairaalassa3,
"Teholla":teholla3,
"Kuolleita":kuolleita3,
"Uusiakuolleita":uusiakuolleita3,
"Toipuneita":toipuneita3
}
df2 = pd.DataFrame(data)
#print(kaikkiatapauksia3)
#print ("Fin wiki data.")
return(df2)
def catch_forecasting_data(toka, last1):
lentoka=len(toka)-2
jono=[]
initu=lentoka
for m in range(0,10):
q=0
for n in range(initu,0, -1):
cc=toka[n]
jono.append(cc)
if (cc<last1):
initu=n-1
if(q==0):
jono.pop()
break
arra00=np.array(jono).astype(int)
arra=arra00[::-1]
return(arra)
-
-
- df=load_country_cases('Finland')
- df2=cut_by_dates(df, paiva1,paiva2)
- countries=["Finland", "Denmark", "Israel", "Sweden"]
- countries=["Finland", "Denmark", "Israel", "France", "Germany", "Italy", "Belgium", "Netherlands", "Portugal", "Austria", "Brazil", "Lithuania", "Mexico", "Argentina", "Latvia"]
countries=["Finland", "Denmark", "Israel", "France", "Germany", "Italy", "Belgium", "Netherlands", "Portugal", "Austria", "Brazil", "Lithuania", "Mexico", "Argentina", "Latvia"]
num_countries=len(countries)
dfs=[]
for n in range(0,num_countries):
#print (countries[n])
dfs.append(load_country_cases(countries[n]))
- for n in range(0,num_countries):
- print(dfs[n])
dada=[]
for n in range(0,num_countries):
df=dfs[n]
#print(df)
taily=df['Daily_Cases']
#print(taily)
taily2=np.array(taily).astype(int)
dada.append(taily2)
- for n in range(0,num_countries):
- print(dada[n])
- last cases num of "Finland" data
eka=dada[0]
numu=30
last1=eka[len(eka)-2]
arras=[]
for n in range(1,num_countries):
toka=dada[n]
arra=catch_forecasting_data(toka, last1)
arras.append(arra)
print(arra)
- quit(-1)
maaras=num_countries-1
parras=[]
maaraz=0
for n in range(0,maaras):
arra=arras[n]
parra=arra[0:numu]
print (parra)
qualik=len(parra)
if (qualik>12):
parras.append(parra)
maaraz=maaraz+1
print ("Precess")
supercache=[]
for m in range(0,numu):
cache=[]
for n in range(0,maaraz):
parra=parras[n]
print (parra)
mmm=parra[m]
cache.append(mmm)
supercache.append(np.array(cache).astype(int))
average00=[]
deviation00=[]
min00=[]
max00=[]
for n in range(0,numu):
average00.append(statistics.mean(supercache[n]))
deviation00.append(statistics.stdev(supercache[n]))
min00.append(np.min(supercache[n]))
max00.append(np.max(supercache[n]))
average1=np.array(average00).astype(int)
deviation1=np.array(deviation00).astype(int)
min1=np.array(min00).astype(int)
max1=np.array(max00).astype(int)
high1=average1+deviation1/2
low1=average1-deviation1/2
- ystdev=statistics.stdev(ycasesminusavgol)
- mean=statistics.mean(ysavgol)
- print (parra)
vormat1='%Y-%m-%d'
today = date.today()
- print("Today's date:", today)
yesterday = datetime.today() - timedelta(days=1)
timedelta1= np.timedelta64(numu,'D')
beginday=today
endday=beginday+timedelta(numu)
days1 = pd.date_range(beginday, endday, periods=numu).to_pydatetime()
- print (parra)
dateformat1 = mdates.DateFormatter('%d.%m')
plt.xticks(fontsize=16)
plt.yticks(fontsize=16, rotation=0)
ax1=plt.gca()
plt.title("Koronavirustapauksia - ennuste muiden maiden datoista", fontsize=18)
ax1.set_xlabel('Pvm', color='g',size=18)
ax1.set_ylabel('Koronatapauksia', color='#800000',size=18)
ax1.xaxis.set_major_formatter(dateformat1)
for n in range(0,maaraz):
plt.plot(days1, parras[n],color='#800000', alpha=0.1 )
plt.plot(days1, average1,color='#F00', linewidth=3)
plt.fill_between(days1, max1, min1, fc='#FF3F3F', alpha=0.25)
for n in range(0,6):
high1=average1+(2*deviation1)/(n+1)
low1=average1-(2*deviation1)/(n+1)
plt.fill_between(days1, high1, low1, fc='#FF3F3F', alpha=n*0.05)
for n in range(0,6):
high1=average1+(deviation1)/(n+1)
low1=average1-(deviation1)/(n+1)
plt.fill_between(days1, high1, low1, fc='#FF1F1F', alpha=n*0.05)
- plt.plot(days1, parras[n],color='#800000', alpha=0.1 )
plt.ylim(0,6000)
plt.show()
- print (last1)
Licensing
edit- You are free:
- to share – to copy, distribute and transmit the work
- to remix – to adapt the work
- Under the following conditions:
- attribution – You must give appropriate credit, provide a link to the license, and indicate if changes were made. You may do so in any reasonable manner, but not in any way that suggests the licensor endorses you or your use.
- share alike – If you remix, transform, or build upon the material, you must distribute your contributions under the same or compatible license as the original.
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 10:14, 10 December 2021 |  | 1,058 × 518 (50 KB) | Merikanto (talk | contribs) | update |
| 07:08, 16 July 2021 |  | 905 × 486 (55 KB) | Merikanto (talk | contribs) | update | |
| 06:33, 25 June 2021 |  | 794 × 454 (108 KB) | Merikanto (talk | contribs) | update | |
| 12:54, 3 June 2021 |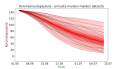  | 904 × 519 (109 KB) | Merikanto (talk | contribs) | Upload | |
| 12:11, 3 June 2021 |  | 916 × 507 (80 KB) | Merikanto (talk | contribs) | Update | |
| 07:56, 3 June 2021 |  | 848 × 526 (38 KB) | Merikanto (talk | contribs) | Update | |
| 13:56, 23 March 2021 |  | 974 × 511 (61 KB) | Merikanto (talk | contribs) | Update | |
| 18:49, 26 February 2021 |  | 841 × 432 (74 KB) | Merikanto (talk | contribs) | ipdate | |
| 10:53, 22 January 2021 | 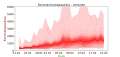 | 910 × 432 (65 KB) | Merikanto (talk | contribs) | Upload | |
| 11:15, 2 January 2021 |  | 1,013 × 512 (65 KB) | Merikanto (talk | contribs) | Update |
You cannot overwrite this file.
File usage on Commons
There are no pages that use this file.
Metadata
This file contains additional information such as Exif metadata which may have been added by the digital camera, scanner, or software program used to create or digitize it. If the file has been modified from its original state, some details such as the timestamp may not fully reflect those of the original file. The timestamp is only as accurate as the clock in the camera, and it may be completely wrong.
| Width | 846.72pt |
|---|---|
| Height | 414.72pt |